Research
Our lab combines bioinformatics, biochemistry, structural biology and molecular biology to explore enzymes that have been sparsely characterized, misidentified or otherwise neglected. Our research highlights include:
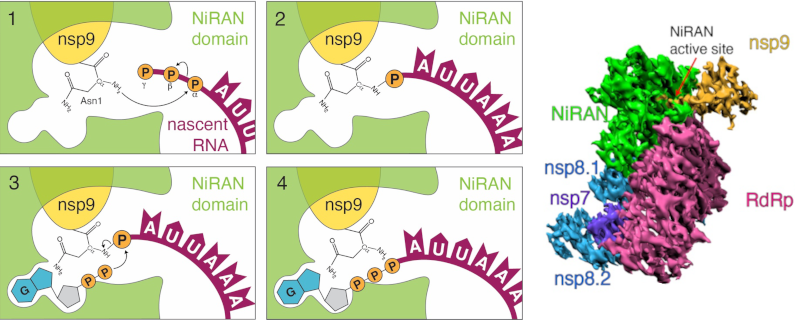
Our elucidation of the mechanism by which SARS-CoV-2 caps its genome. Capping requires RNAylation, in which a virally encoded pseudokinase catalyzes the transfer of RNA to the N-terminal amino acid of another SARS-CoV-2 protein as an intermediate step.
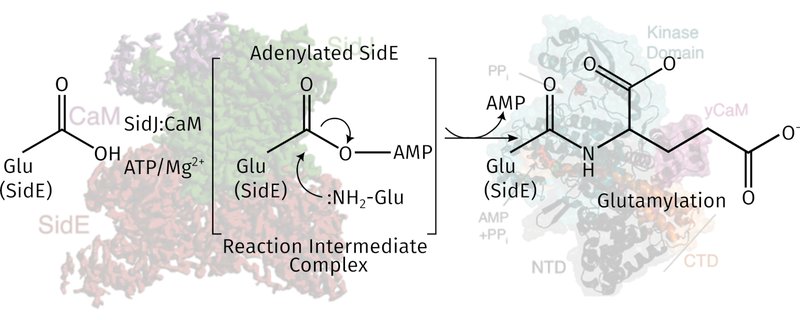
Our work on the legionella metaeffector SidJ, which we discovered to be a polyglutamylator of the SidE family of ubiquitin ligases. We pinned down its mechanism using a series of biochemical experiments and by solving multiple structures of the complex.
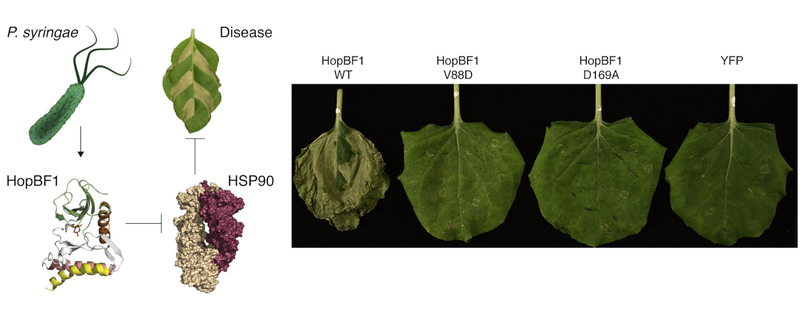
Our identification of the HopBF1 bacterial effector as a betrayer of Hsp90, in which HopBF1 poses as a pseudoclient of the chaperone in order to deactivate it.
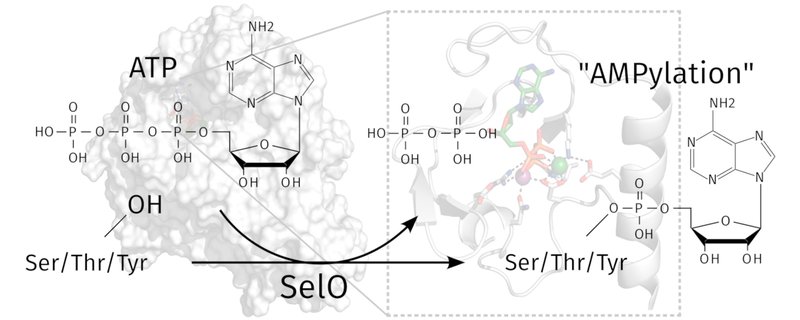
Our study of the protein SelO, which adopts a protein kinase fold and functions by catalyzing the transfer of AMP to its substrates.